  
This is most commonly achieved by first estimating a phylogeny with branch lengths (in units of substitutions per site), then adjusting these branch lengths so that they are proportional to time (often called a chronogram or time tree). Regardless of methodology, molecular dating relies on two processes: (1) estimating substitution rates among sequences and (2) calibrating substitution rates with independent evidence to convert estimated genetic distances (usually in units of substitutions per site) to absolute ages. These methods have become increasingly more statistical and have a strong foundation in molecular evolution and systematics. These are a general class of methods often referred to as “relaxed” clocks. With this renewed interest, there has also been great effort made to modify the assumptions of a “strict” clock to account for rate variation among lineages. Despite these concerns, the use of molecular clock methods has seen a renewed interest in the past fifteen years. Over the years, the ability to estimate divergence times among species in this manor has been met with great skepticism. This theory also provided an important null model of molecular evolution, but was not without its critics. This theory proposed that most of the substitutions that we observe in molecular data (and the variation we see within species at the molecular level) is due to the fixation of these changes that are neutral or nearly neutral with respect to selection. Kimura’s neutral theory of molecular evolution provided an explanation of why macromolecules might be evolving in a clock-like fashion. 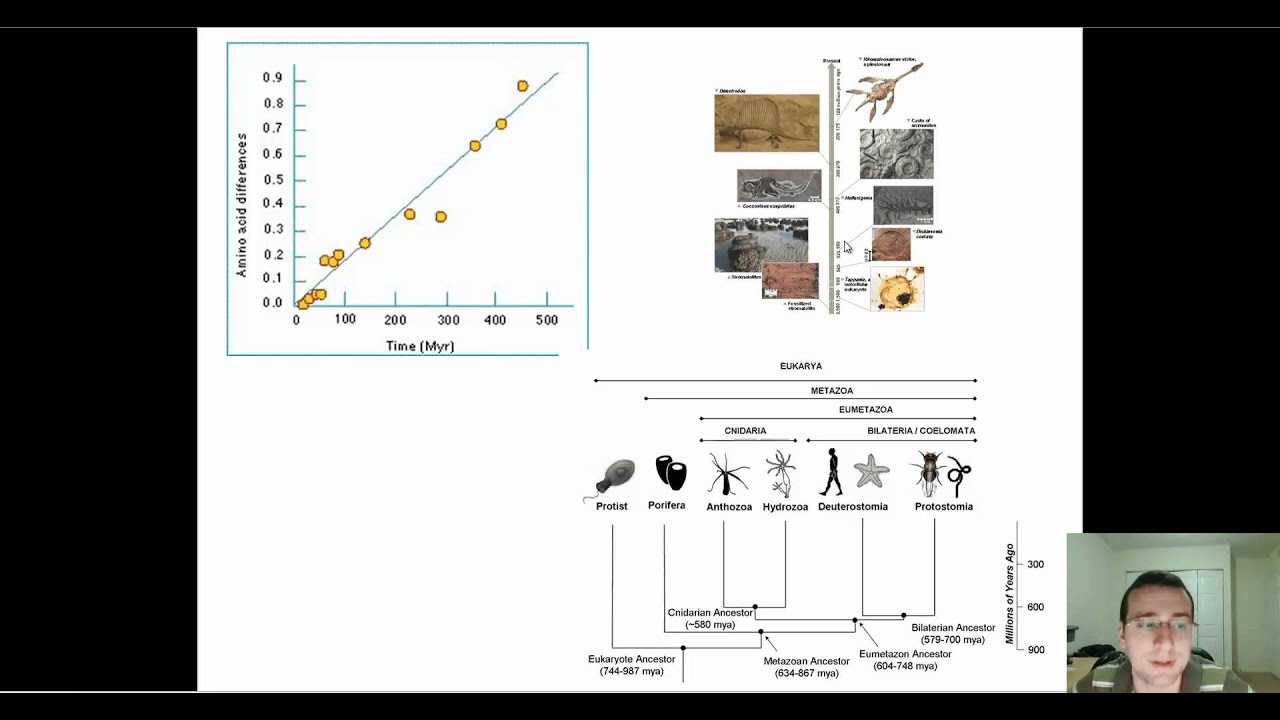
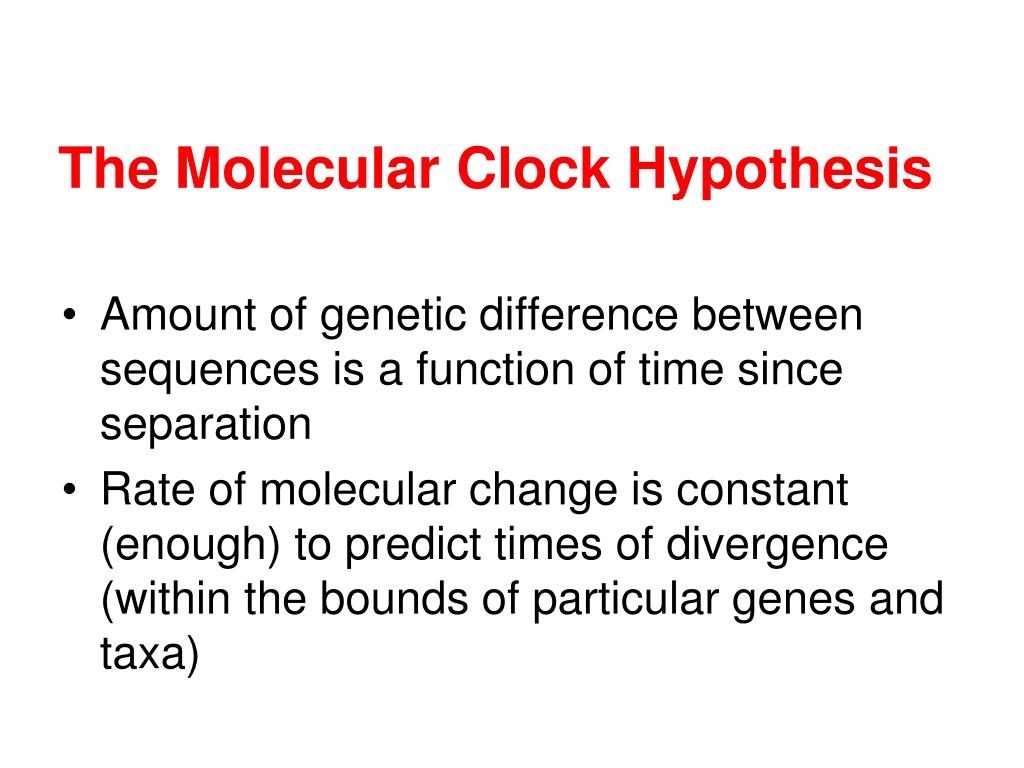
Although Zuckerkandl and Pauling provided evidence for a linear relationship between the accumulation of amino acid differences and evolutionary time, they did not provide an explanation for “why” they observed this pattern. Because of this, the absolute time since the last common ancestor for species must then be calculated by calibrations based on paleontological evidence. One major issue of using sequence data to infer absolute divergence times is how to disentangle time from evolutionary rates. In addition, they suggested that this uniform rate of a specific protein would be approximately constant, not just over evolutionary time, but also across different lineages or taxonomic groups. That is, differences between sequences would accumulate in a linear fashion. The authors proposed that amino acid differences in a protein should accumulate at, more or less, a uniform rate across different species. The concept of a “molecular clock” was originally suggested by Emile Zuckerkandl and Linus Pauling based on their observations of amino acid substitutions between hemoglobin sequences. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
